Micro and Macro Brain Circuits
One of the focus of the lab is to determine the relationship between the cellular/tissue architecture and brain functions with emphasis on: a) the role of neuronal morphology in the mechanisms for information processing, b) the optimal ways to wire the brain “from microcircuits to global graph theory”, and c) the relationship between neuronal excitability (spiking – outputs, M(S)UA) and presynaptic input reflections (local field potentials, LFP). We previously worked on using multi-photon calcium imaging and whole-cell patch clamp in acute slices of the cerebral cortex of Wistar rats to formulate realistic biophysical models for the genesis of LFP/M(S)UA by large layer V pyramidal cells. In collaboration with NeuroNexus, a high-resolution 3D silicon-probe that comprises 128 microelectrodes arranged within 1 mm3 cortical volume (“Custom Design Portfolio”, Fig. 1) was developed. By the use of this groundbreaking technology, the group has been able to study neuronal excitability not only at the level of neuronal masses but also that related to the activity of single neurons. A major interest of the group is the characterization of the coding/decoding strategies in the somatosensory cortices of rodents (e.g. complex representation of whiskers in the barrel field). We are also using this ground-breaking technique to characterize ion electro-diffusion process in the neocortex. Electroencephalographic (EEG) techniques in rodents (EEG minicap) and humans (EasyCap) have been used to study neuronal activity at the system level from healthy and pathological brains. By using geometrical and functional descriptors from fluorescent images, we have recently developed multi-compartmental models of different neuron types and used them to simulate the activity of neuronal masses through parallel MATLAB code running in clusters of personal computers. These biophysical models of the cellular/tissue architecture have been employed in the laboratory to fuse and interpret observable data from different neuroimaging modalities (e.g. Fig. 2, EEG and fMRI data in a visual stimulation protocol).
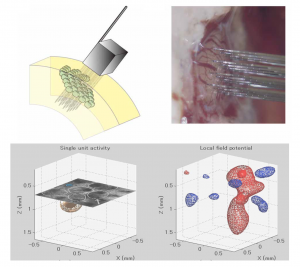
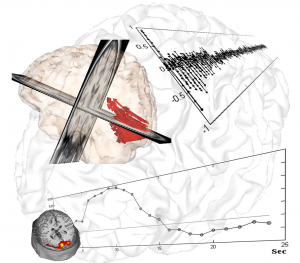
Particular ongoing projects are:
- Cortical spreading depression mechanisms (Arash Moshkforoush, Daniel Rivera, Diana Borrego, Pedro Valdes-Hernandez) – In collaboration with Dr. Yoichiro Mori (University of Minnesota)
- Brain networks in epilepsy (Pedro Valdes-Hernandez, Divya Teltumbade, Patrick Henry, Elena Atienza, Fernando Gonzalez) – In collaboration with Dr. Byron Bernal (Nicklaus Children Hospital)
- Neurosensing project in epilepsy (Carolina Moncion) – In collaboration with Dr. John Volakis (ECE-FIU)
Neurovascular Coupling
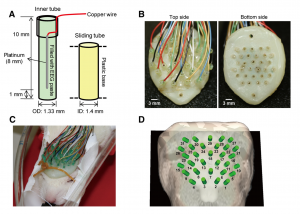
We have developed models for fusing EEG and fMRI multimodal data based on the major principles for the genesis/diffusion of nitric oxide, a well-studied phasic pathway connecting neurons and vessels in the cerebral cortex. Such models were used to interpret multimodal experimental data from humans undergoing visual, somatosensory and motor tasks. These models have been additionally useful to understand the large-scale functional connectivity profiles in the human brain from combining hemodynamic/metabolic-based imaging modalities and electrophysiological data. Recently, we have developed a novel minicap for whole head EEG recording in rodents, which is currently US patented (Fig. 1). Based on this technique, members of the team record EEG signal concurrently with local blood flow activity (Laser Doppler flowmeter, Perimed Inc, US) and O2/NO tissue metabolism (Amperometric microsensor, Unisense, DA) under a variety of pharmacological manipulations in vivo. To perform data fusion and proper stimulation protocols in Wistar rats, we have developed a brain probabilistic atlas for this particular species and introduced novel methods for image co-registration. We have combined this technique with fMRI BOLD effect to study neuro-vascular/metabolic coupling in the neocortex of rodents. Of particular interest is the comprehension of the role played by the protoplasmic astrocytes in: a) the modulation of neuronal activity, b) signaling-based regulation of local brain metabolism, and c) vessel controllability, both in normal and pathological brains. Also, we have developed a novel technique based on the combination of multi-photon calcium imaging and multi-side intracranial recordings in vivo. To study hemodynamic and metabolic abnormalities in the epileptic tissues, this technique is being now extended to other optical imaging modalities (e.g. diffuse reflectance spectroscopy). With the help of optogenetic methods and fluorescent-loading protocols, the latter approach allows our team to quantify those astrocytic-derived factors with actions into both the neuronal networks and the cerebral microcirculation. In the past, we had postulated that dysfunctions in the astrocytic networks might be an important component in the physiopathology of Alzheimer disease. The introduction of biophysical models for the genesis and propagation of intercellular signaling has been crucial to evaluate the imprints left by certain brain disorders in functional neuroimaging.
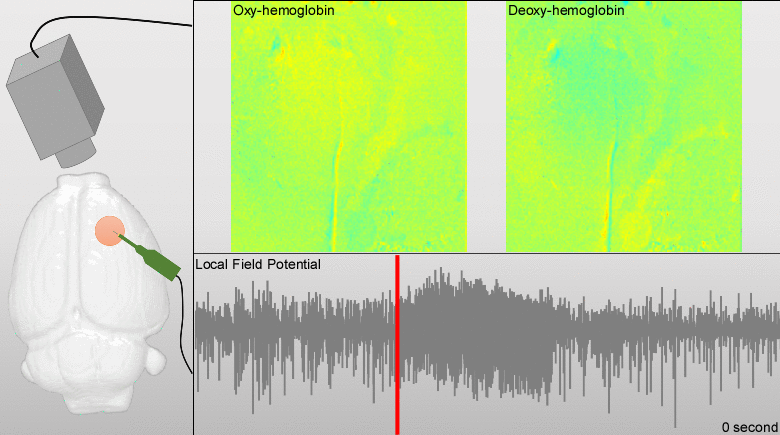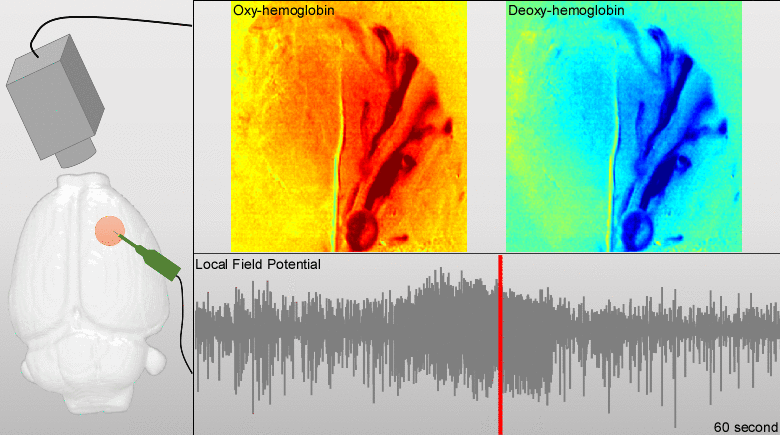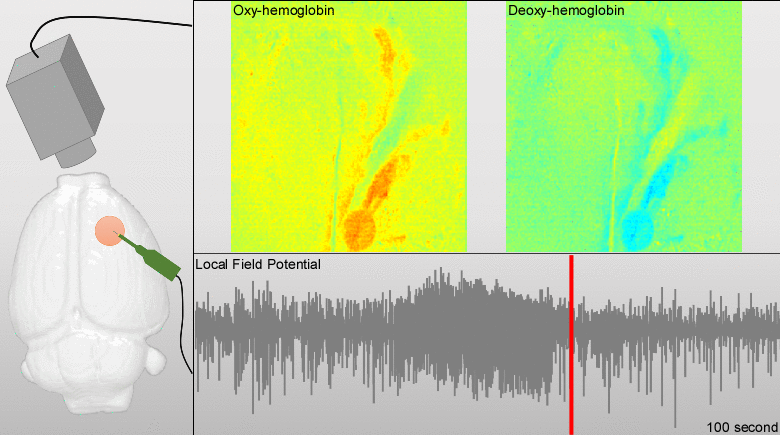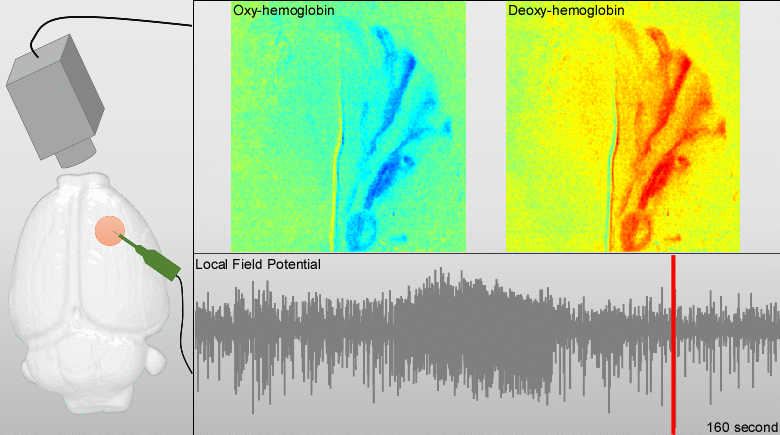
Particular ongoing projects are:
- Negative BOLD effect in normal and pathological conditions (Pedro Valdes-Hernandez) – In collaboration with Dr. Fahmeed Hyder (Yale University) and Dr. Wei-Chiang Lin (BME-FIU)
- Neurovascular mechanisms in cortical spreading depression (Arash Moshkforoush) – In collaboration with Dr. Nikolaos Tsoukias (BME-FIU)
- EEG-fNIRs fusion to study neurovascular coupling in hypertension (Ernest Mares, Carolina Moncion) – In collaboration with Dr. Anuradha Godavarty (BME-FIU)
Neuro-Inflammatory Processes
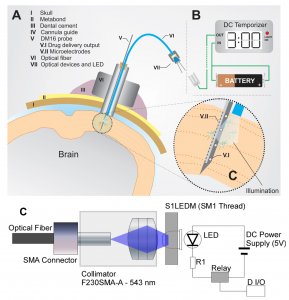
In the last five years, we have focused on understanding astrocytic calcium signaling in both normal and pathological brains. Being a pioneer in combining intracranial recordings and multi-photon laser scanning microscopy in vivo to access the activity of networks of neurons and astrocytes, we have also used similar techniques to study dysregulations in the astrocytic calcium activity caused by deposition of β-amyloid plaques. It is worth to hypothesize that endogenous/induced cyto-architectonic changes (e.g., FCD lesions, strokes, Ab plaques) might create progressively astrogliosis evoked by undergoing inflammatory processes, and consequently, cause local dysregulations in the astrocytic calcium signaling. The main objective of this research line is to determine the major causes of both alterations in astrocytic calcium signaling and the associated inflammatory signaling associated with brain lesions, as well as to develop an optogenetic method to prevent these alterations (Fig. 1), and hence, the possible development of epileptic seizures. A non-invasive control with light stimulation of such dysregulations in the astrocytic calcium activity is expected to interrupt the mechanisms for the novo-synthesis of pro-inflammatory signals.
Particular ongoing projects are:
- Epileptogenesis in focal cortical dysplasia (Lakshmini Balachandar) – In collaboration with Dr. Jeremy Chamber (Cellular Biology and Pharmacology, FIU